The Enigma of Drug Release
How pharmaceutical release modeling mirrors WWII cryptography, and how digiM's ivisLab and dissoLab replace blind assumptions with real 3D microstructure data.

Growing up, I was obsessed with cryptography. The perfect asymmetry of a cipher, simple to scramble, nearly impossible to reverse.
Take the WWII Enigma machine. To crack it, Alan Turing, the father of modern computing and his team built a brilliantly sophisticated computational device. But here’s the catch: the machine alone wasn't enough. To solve the puzzle, the machine needed a "crib", a known snippet of text, like a daily weather report, hidden within the encrypted message. Knowing just a few exact starting words anchored the math, turning an impossible puzzle into a solved one.
What does WWII cryptography have to do with pharmaceuticals? A surprising amount.
Predicting how a complex drug releases a drug into the body, is essentially cracking a microscopic cipher.
The Problem: A Brilliant Machine Without a Crib
Conventionally, pharmaceutical modeling teams build their own "Turing machines": highly sophisticated physical and chemical release models. They are brilliant mathematical solvers.
But there's a problem. To make the math work, scientists assume their starting conditions. They pretend drug particles are perfect spheres or perfectly blended. They are running a brilliant machine without a crib. Because a real system is messy and chaotic, the math breaks down, forcing modelers to rely on unphysical "fudge factors" to make their predictions match reality. And because these fudge factors exist to correct initial conditions that never matched reality there is no guarantee they will work when the samples change.
The Solution: Finding our crib
What if we stopped guessing the starting conditions and just looked? That’s the exact philosophy behind digiM’s ivisLab and dissoLab
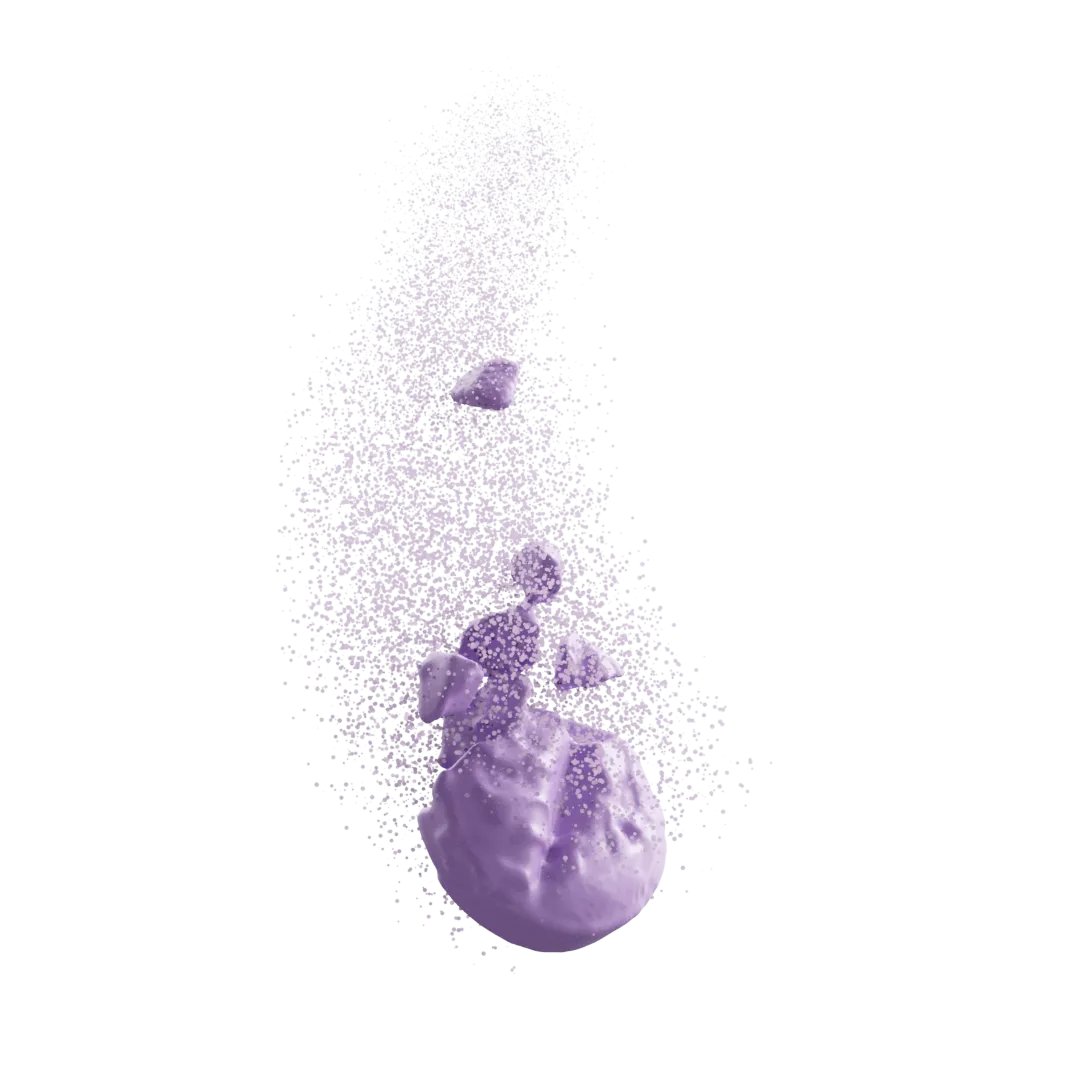
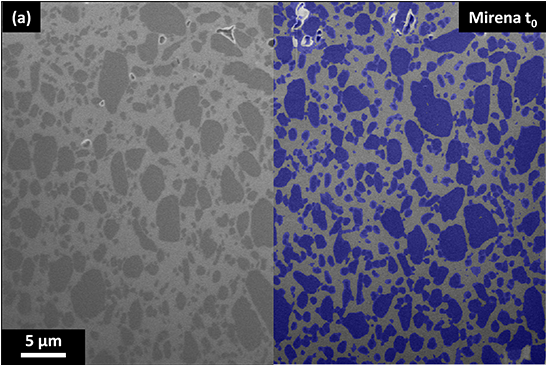
- The Crib (Imaging): Instead of assuming internal structure, we use high-resolution 3D X-ray imaging and AI image analysis to map the real, as-manufactured microstructure. By capturing the exact location of the pores, polymer, and active ingredients (API), ivisLab gives us undeniable initial conditions, our cryptographic crib.
- The Machine (dissoLab + ivisLab): We then feed that exact 3D reality directly into our solver. Because our physical and chemical models run on actual measured geometry instead of blind assumptions, the math just works. The simulation calculates how fluid and API move through the real, tortuous pore network naturally, no fudge factors required.
By pairing a brilliant solver with perfect initial conditions, the blind guesswork vanishes. We stop forcing math and start predicting reality.
Transform Your Program with Microstructure Science
Get started with a drug product digital twin.


















.jpeg)