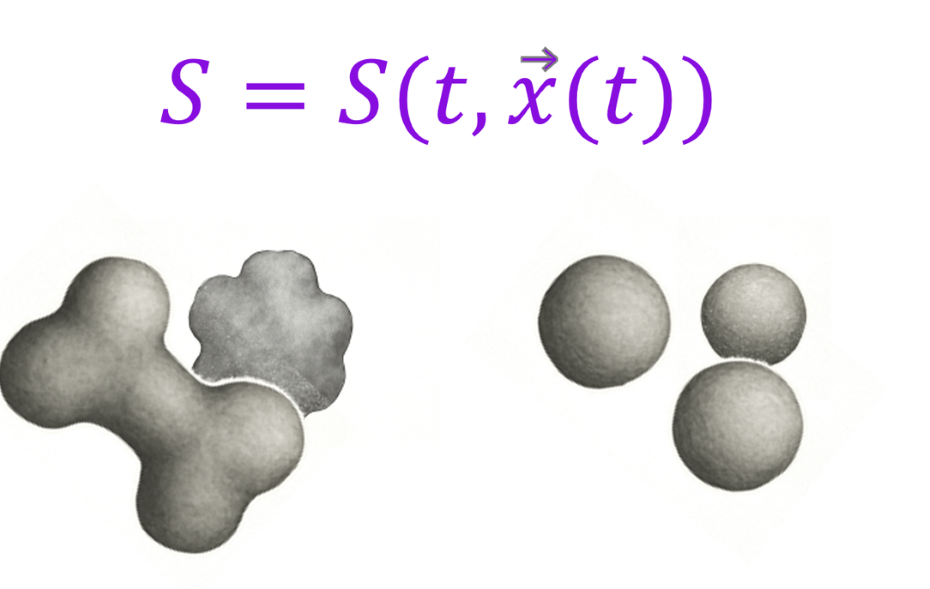Dissolution prediction from images is neither magic, nor regression
Latest updates and common Q&A about image-based dissolution prediction

Since the release of dissoLab in January, it generated a lot of interest across different departments in the pharmaceutical industry, and across different industries including regulatory agencies. Amid the enthusiasm, a few common questions often bring a smile, or even a laugh, as they are “dissolved” in the discussion.
Dissolution modeling has been done for decades. So what’s new with dissoLab?

dissoLab tracks the full surface area evolution throughout the dissolution process, based on actual sample images, on a 3D per-pixel basis. That may sound like a mouthful. Mathematically, dissoLab handles surface area as a temporal and spatial function. Conventional dissolution models simplify the particles into spherical geometry. They do not consider surface roughness, aspect ratio, and complex morphological behaviors such as aggregation, intra-particle porosity, wall breakage, and particle split, all of which were considered by dissoLab. While conventional models apply Noyes-Whitney globally, dissoLab applies Noyes-Whitney locally on a per-pixel basis. In addition to S - surface area being a temporal and spatial function, D - drug diffusivity, h - boundary layer thickness, Cs - drug solubility, and V - media volume, are (or will be in future versions) all modeled locally on a per-pixel basis as a temporal and spatial function.
Some Common Questions
How can you possibly handle all these different, complex, and often unpredictable morphological behaviors?
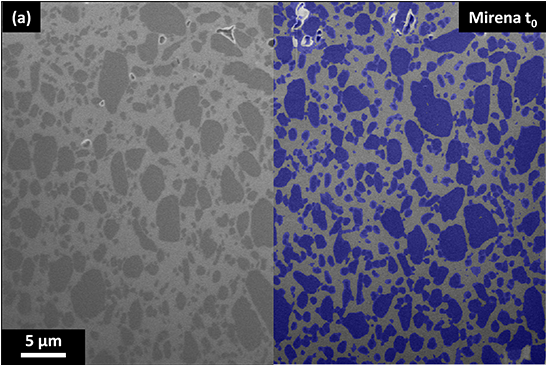
dissoLab handles these behaviors implicitly by applying morphological operations on the imaging data. After ensuring that the particles captured in the images are representative, their morphological behavior can be tracked precisely using pixel erosions.
How can you computationally track many thousands of particles?
Well, why should that matter if the results are fast and valid? All jokes aside, dissoLab uses highly optimized 3D morphological operators with GPU accelerated structure synthesis. While it might appear unrealistically fast compared to MATLAB or Python research codes, dissoLab, along with other digiM I2S software algorithms, are engineered for industrial-scale performance..
How long does dissoLab run? What are the most time consuming steps?
A typical dissoLab run takes between a few seconds up to a couple of hours. The most time consuming case is when the user inputs a 2D image, which needs to be representatively synthesized into 3D, taking a couple of hours. dissoLab requires input images to be segmented. Depending on the quality of the images and complexity of the features, segmentation can take a few minutes (using digiM seg2D software) or a few hours. digiM can help with segmentation through our contract research team.
How do you ensure representativeness?
This is a great question, while the answer goes beyond the scope of dissoLab. The rationale is simple – if an image of a spherical particle is provided, dissoLab cannot predict the dissolution of a sample with a cube particle only using that input. Garbage in, garbage out, like in any modeling software.
With that said, dissoLab provides a suite of tools that can enhance representativeness from limited input data.
• When the input is a 2D image with a small number of particles, dissoLab employs a sGAN structure synthesis deep learning tool to
Transform Your Program with Microstructure Science
Get started with a drug product digital twin.